Research Techniques
DNA Curtains
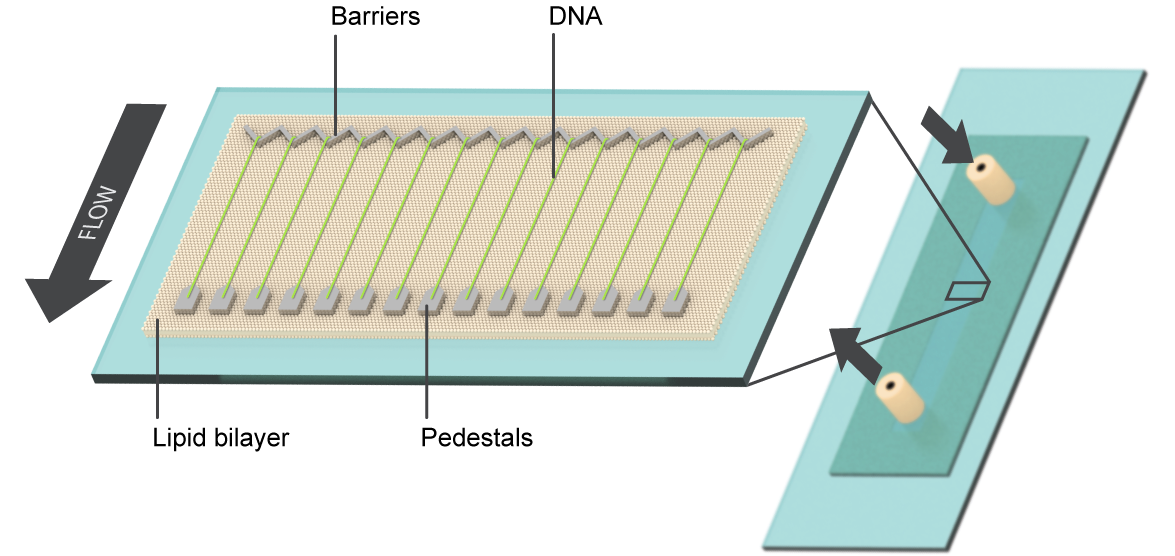
This single-molecule optical microscopy technique is used to study fundamental interactions between proteins and nucleic acids, using a non-traditional combination of biochemistry, physics, and nano-scale technology. It allows us to observe individual protein molecules and protein complexes as they interact with their DNA substrates. The goal is to reveal which proteins bind to DNA, where they bind, how they move, and how they influence other components of the system in real-time, on a single-reaction scale.
Our lab developed the DNA curtain technique to overcome limitations on a single-reaction scale in other TIRFM-based biochemistry. The technology uses fluid lipid bilayers to render surfaces inert to biological molecules, and micro- and nano-scale materials engineering to construct parallel arrays containing individual DNA molecules with user-defined positions, orientations, tensions, and topologies. These DNA arrays allow for parallel processing of hundreds to thousands of individual protein-nucleic acid interactions in a single TIRFM experiment and serve as powerful tool for single-molecule research.
Single-particle cryo-electron microscopy
Cryo-electron microscopy (cryo-EM) is a modern structural technique that uses images taken on an electron microscope to reconstruct the 3D structure of proteins and other biological macromolecules. By plunge-freezing samples in liquid ethane, samples freeze without forming ice crystals in a process called vitrification, which preserves molecules in their native state and protects them from being damaged by the electron microscope. Samples are then imaged on an electron microscope, where 2D images are taken of particles in many different orientations that can then be reconstructed into a 3D image.
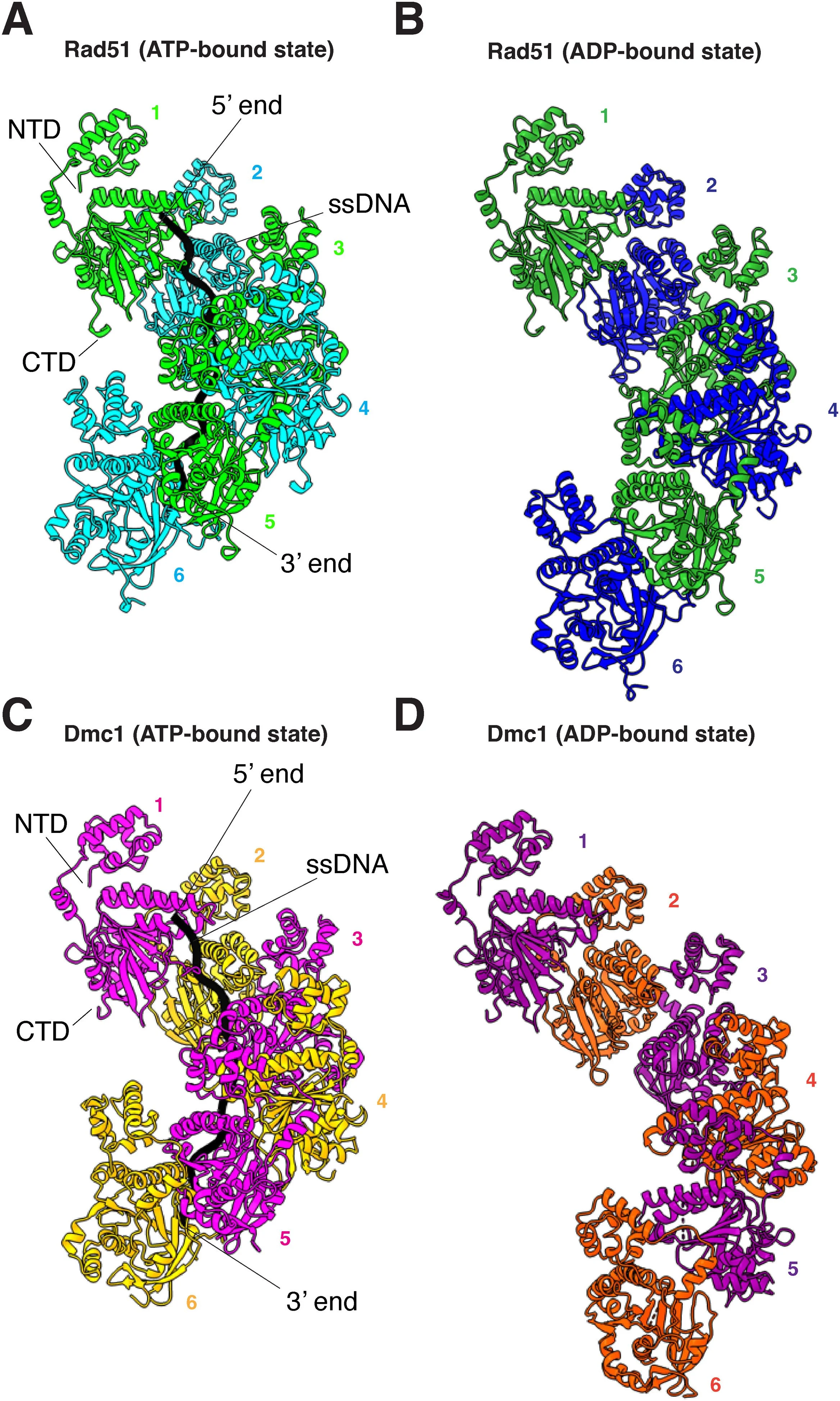
Our lab uses this technique to understand protein-protein and protein-nucleic acid interactions that occur during the process of homologous recombination. Some of our recent work involving cryo-EM has allowed us to understand the structural transitions that occur within yeast Rad51 & Dmc1 filaments during ATP hydrolysis and how the yeast proteins Rad54 & Hed1 interact with Rad51 filaments.
Recent work involving cryo-EM:
Shin Y, Kim SY, Greene EC. ATP hydrolysis-driven structural transitions within the S. cerevisiae Rad51 and Dmc1 nucleoprotein filaments. J Biol Chem. 2025 Jul 26:110528. doi: 10.1016/j.jbc.2025.110528. Epub ahead of print. PMID: 40721016.
Shin Y, Petassi MT, Jessop AM, Kim SY, Matei R, Morse K, Raina VB, Roy U, Greene EC. Structural basis for Rad54- and Hed1-mediated regulation of Rad51 during the transition from mitotic to meiotic recombination. bioRxiv [Preprint]. 2025 Mar 26:2025.03.26.645561. doi: 10.1101/2025.03.26.645561. PMID: 40196570; PMCID: PMC11974805.
Deep Mutational Scanning (DMS)
DMS, or deep mutagenesis, is high-throughput technique used to analyze the function of single-amino-acid-substitution mutations in proteins. This technique relies on random mutation of the codon of the amino acid of interest to every other codon, thereby generating a vast library of mutations. This library is introduced into a model organism and subjected to selective pressure. The frequency of each amino acid residue is identified before and after selective pressure using next-gen sequencing, which allows us to observe the relative impact of the mutations to the selected phenotype.